AUTOMATED SYSTEMS FOR BACTERIAL IDENTIFICATION AND ANTIBIOGRAM: VITEK 2 AUTOMATED COMPACT SYSTEM & MALDI-TOF
There are several commercially available automated systems or computerized instruments and/or equipments that aid in the clinical/laboratory diagnosis of infectious diseases in the hospital especially as it relates to prompt and accurate identification of microbes. These automated systems also aid in carrying out antimicrobial susceptibility test (antibiogram) on identified clinical pathogens so that antimicrobial therapy can be properly guided.
The VITEK 2 automated compact system (BioMérieux-Vitek) is one of such advanced expert systems that could be used in clinical microbiology laboratories for the prompt identification of microbial pathogens and for the determination of the antimicrobial susceptibility of already identified microbes (inclusive of bacteria and fungi). The VITEK 2 automated system is an automated high-throughput microbial identification system that uses an innovative mass spectrometry (MS) technology such as the Matrix Assisted Laser Desorption Ionization Time-of-Flight (MALDI-TOF) to provide pathogen identification results and antimicrobial susceptibility test (AST) results in minutes.
It detects metabolic changes by fluorescence-based methods which facilitate the identification of Gram-negative bacteria, Gram positive bacteria and pathogenic fungi from clinically-relevant specimens within some few hours. The VITEK 2 system monitors the kinetics of bacterial and/or fungal growth and calculates the minimum inhibitory concentrations of (MICs) of antimicrobial agents or antibiotics using a unique algorithm with taxonomically updated databases of microbial strains and AST profiles.
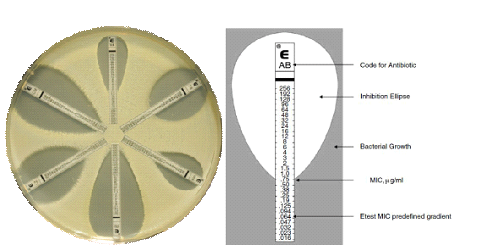
The VITEK system includes an Advanced Expert System (AES) that analyzes minimum inhibitory concentration (MIC) patterns of antimicrobial agents or antibiotics; and this automated system also detects phenotypes for most microbial organisms including Gram positive bacteria, Gram negative bacteria and yeast-like organisms (Figure 1). The VITEK systems provide rapid laboratory test results especially as it is related to bacterial identification and antibiogram. Rapid laboratory results allow clinicians/physicians to discontinue blind treatment or empiric antimicrobial therapy (which allow resistant pathogens to evolve and spread), and thus prescribe targeted antimicrobial therapy that would result into improved patient outcomes and enhanced antibiotic stewardship in this era of skyrocketing antibiotic resistance.

On a daily basis, medical doctors use results from the identification of pathogens from clinical specimens and antibiotic susceptibility test (AST) results to determine the antibiotic treatment that is most appropriate for a particular patient or disease in order to achieve best patient care. Also, the results from the AST are used to monitor changes in bacterial resistance to antibiotics in order to detect and halt any possible outbreak of disease due to resistant pathogens. Bacterial identification and antibiotic susceptibility testing play an essential role in patient care and the control of antibiotic resistance.
These important laboratory procedures indicate the aetiology of the disease and also the likely antibiotics most suitable to cure or treat the infection or disease. AST also help physicians to avoid the unnecessary prescription of antibiotics – which help to reduce healthcare costs and prevent the evolution of drug resistant organisms. However, these important clinical microbiology procedures (i.e. identification of pathogenic organisms and AST) are usually delayed in some hospitals because they lack the equipments and or instruments required to carry out these techniques promptly and deliver accurate results under the shortest possible time.
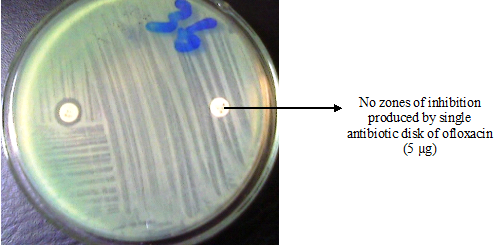
The VITEK 2 system or platform is an AES that offers automated bacterial identification and AST so that clinical results as it pertains to AST and bacterial identification could be obtained within minutes or hours of processing the sample (Figure 2). In the conventional microbiological culture technique of bacterial identification and AST, clinical results pertaining to microbial identification and AST usually takes days to get; and this impact negatively on the patient’s prognosis since blind treatment may be initiated until the test results from the clinical microbiology laboratory is finally obtained.
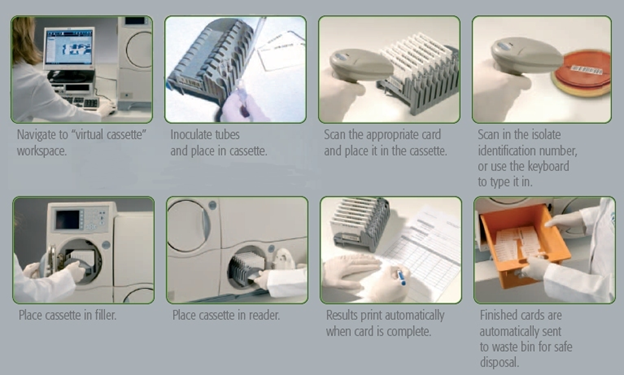

Each VITEK 2 identification card has several rows of wells comprising of 8 rows of 8 wells that is made up of 64 wells per card (8 wells X 8 wells = 64 wells). The wells of the identification card contain different dehydrated media required for different biochemical tests targeted at identifying via biochemical reactions different microbial strains. These individual biochemical tests are meant for the identification of Gram negative bacteria, Gram positive bacteria and fungi (yeast cells). A capillary tube (which sucks the suspension of bacteria to be identified and dispenses into all the wells) is fixed to each VITEK 2 identification card.
Microbial (bacterial or fungal) growth is spurred by the hydration of the dehydrated media in the in the card with a suspension liquid (e.g. distilled water). The cards are incubated in the card reader / incubation chamber of the VITEK 2 compact system; and the colour changes in all the wells are recorded automatically in the VITEK 2 compact system. The results of the color changes go to a computer system that is attached to the VITEK 2 compact system. The computer system has an inbuilt updated database of different bacterial and fungal organisms as per their unique biochemical reactions. The computer automatically compares the results with those available in its library for different bacteria and/or fungi, and finally gives the name of the bacteria with a definite probability.
For identification of the organism, the given bacterium or fungal organism, grown as isolated colony on a culture media plate or as pure culture grown on a slant are taken; and a loopful of the bacteria is transferred aseptically into sterile saline solution in a test tube and a suspension of the organism is made. The suspension should contain a prescribed density of bacteria, as determined by a densitometer. The test tube is fixed to the cassette and a card is fixed near it, such that the tip of the suction capillary tube of the card remains deeply submerged in the suspension. The number of test tubes and cards fixed to each of the cassette can be increased depending on the number of bacteria or fungi to be identified by the VITEK 2 identification cards. The cassette is put in the vacuum chamber of the system as shown in the illustration.
A high vacuum is created inside the chamber, which forces the bacteria suspension to be sucked into the capillary tubes and dispensed into the wells of the cards. The cassette is taken out and put inside the incubation and analysis chamber where the capillary tubes are cut and the cut ends sealed automatically. The incubation process is allowed to run at a prescribed temperature for a prescribed period of time, which is programmed by the control panel on the VITEK 2 compact system. During incubation, which is usually in every 15 minutes, each VITEK 2 identification card automatically goes to the colour reader, which reads the colour changes in the wells and records them. The recorded results go to the computer, which automatically compares them with those, available in its library for different bacteria. Finally, it gives the names of the bacteria with definite probabilities. The used cards fall into the waste disposal chamber (waste collection bin) of the system for removal and final disposal after sterilization.
References
Arora D.R (2004). Quality assurance in microbiology. Indian J Med Microbiol, 22:81-86.
Ashutosh Kar (2008). Pharmaceutical Microbiology, 1st edition. New Age International Publishers: New Delhi, India.
Barenfanger J, Drakel C and Kacich (1999). Clinical and Financial Benefits of Rapid Bacterial Identification and Antimicrobial Susceptibility Testing. Journal of Clinical Microbiology, 37(5):1415-1418.
Denyer S.P., Hodges N.A and Gorman S.P (2004). Pharmaceutical Microbiology. 7th ed. Blackwell Publishing Company, USA.
Doern G, Brueggemannn A.B, Perla R, Daly D, Halkias D, Jones R.N, Saubolle M.A (1997). Multicenter laboratory evaluation of the bioMerieux Vitek antimicrobial susceptibility testing system with 11 antimicrobial agents versus members of the family Enterobacteriaceae and Pseudomonas aeruginosa. J Clin Microbiol, 35:2115–2119.
Doern G, Vautour R, Gaudet M, Levy B (1994). Clinical impact of rapid in vitro susceptibility testing and bacterial identification. J Clin Microbiol, 32:1757–1762.
Doern G.V (1995). Susceptibility tests of fastidious bacteria. Manual of Clinical Microbiology, 6th edition, Murray P.R, Baron E.J, Pfaller M.A, Tenover F.C, Yolken R, American Society for Microbiology, Washington DC, Pp. 1342-1349.
Funke G, Monnet D, deBernardis C, von Graevenitz A, Freney J (1998). Evaluation of the VITEK 2 system for rapid identification of medically relevant Gram-negative rods. J Clin Microbiol, 36:1948–1952.
Garcia L.S (2010). Clinical Microbiology Procedures Handbook. Third edition. American Society of Microbiology Press, USA.
Hart C.A (1998). Antibiotic Resistance: an increasing problem? BMJ, 316:1255-1256.
Livermore D.M, Winstanley T.B, Shannon K.P (2001). Interpretative reading: recognizing the unusual and inferring resistance mechanisms from resistance phenotypes.
J Antimicrob Chemother, 48 Suppl 1:87-102.
Madigan M.T., Martinko J.M., Dunlap P.V and Clark D.P (2009). Brock Biology of Microorganisms, 12th edition. Pearson Benjamin Cummings Inc, USA.
Mahon C. R, Lehman D.C and Manuselis G (2011). Textbook of Diagnostic Microbiology. Fourth edition. Saunders Publishers, USA.
National Committee for Clinical Laboratory Standards. Performance Standards for antimicrobial susceptibility testing. 8th Informational Supplement. M100 S12. National Committee for Clinical Laboratory Standards, 2002. Villanova, Pa.
Washington J.A (1993). Rapid antimicrobial susceptibility testing: technical and clinical considerations. Clin Microbiol Newsl, 15:153–155.
Discover more from Microbiology Class
Subscribe to get the latest posts sent to your email.