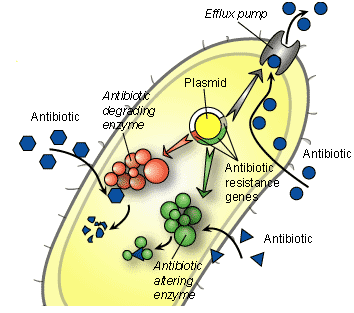
Antibiotic resistant bacteria owe their drug insensitivity and ingenuity in developing resistance against our therapeutic regimens to resistance genes which they harbor or possess. These resistance genes can either be inherently (naturally) part of the organism’s physiology or it can as well be acquired from other resistant organisms in their environment.
It is these resistance genes that resistant bacteria transfer to non-resistant (susceptible) strains of microbes, thus compounding the problem of antibiotic resistance. Some of the major mechanisms via which microbes acquire resistance genes either from their environment or from other organisms include:
- Conjugation
- Transformation
- Transduction
CONJUGATION
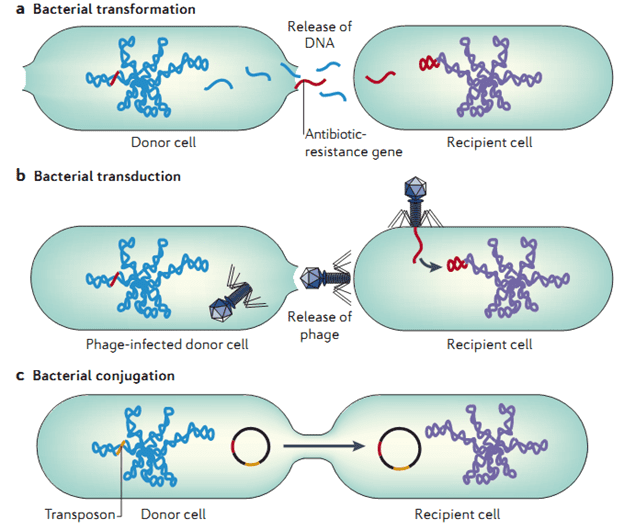
Conjugation is the form of gene transfer and recombination in bacteria through which genetic materials (DNA) are transferred from one bacterium to another through a direct cell – to – cell contact (Figure 1). It occurs by direct contact between two bacteria; and this contact leads to the transmission of genetically important materials such as plasmids amongst bacterial population. Conjugation is the most important genetic transfer mechanism by which bacteria transfer their antibiotic resistance genes to susceptible bacteria.
Conjugation is mediated by a particular kind of circular DNA called a plasmid, which replicates independently of the chromosome. Plasmids are extrachromosomal DNA molecules that are capable of independent replication; and they are not required for the growth and replication of the host cell. Many plasmids carry genes that confer resistance to antibiotics.
When two bacterial cells are in close proximity to each other, a hollow bridge like-structure known as the “pilus” forms between the two cells; this allows a copy of the plasmid as it is duplicated to be transferred from one bacterium to another bacterium (known as the recipient bacterial cell). This process called conjugation enables a susceptible bacterium to acquire resistance genes to a particular antibiotic from another organism which harbours resistant genes.
TRANSDUCTION
Transduction is the transfer of genetic material between bacteria by bacteriophages or phages. Bacteriophages are bacterial viruses (Figure 1). A phage or bacteriophage is a virus that uses bacteria as its host. In transduction, antibiotic resistance genes are incorporated into a phage capsule which is later injected into another bacterium.
In the process of transduction, bacterial DNA is transferred from one bacterium to another inside a virus that infects bacteria. These viruses that infect bacteria are called bacteriophages or phages. When a phage infects a bacterium, it essentially takes over the genetic process of the bacteria to produce more phages. During this process, bacterial DNA may inadvertently be incorporated into the new phage DNA.
Upon bacterial death and lyses or breaking apart, these new phage goes on to infect other bacteria in the environment. This brings along genes from previously infected bacterium into the recipient bacterium. These genes might contain advantageous genes such as antibiotic resistance genes, which will leave the recipient bacterium resistant to a particular antimicrobial agent or antibiotic.
TRANSFORMATION
Transformation is a mechanism of genetic transfer in bacteria in which a piece of free DNA (genetic material) is taken up by a bacterium and integrated into the recipient genome (Figure 1). During this process of transformation, genes are transferred from one bacterium to another as “naked” DNA.
When bacterial cells die and break apart through cell lysis, DNA can be released into the surrounding environment. Other bacteria in close proximity can scavenge this free floating DNA and incorporate them into their own DNA.
This incorporated DNA can contain advantageous genes such as antibiotic resistance genes and benefit recipient bacterial cells – which resultantly become resistant to the antimicrobial agent or antibiotic the resistant gene mediates.
References
Arora D.R (2004). Quality assurance in microbiology. Indian J Med Microbiol, 22:81-86.
Ashutosh Kar (2008). Pharmaceutical Microbiology, 1st edition. New Age International Publishers: New Delhi, India.
Axelsen P.H (2002). Essentials of antimicrobial pharmacology. Humana Press, Totowa, New Jersey, USA. Al-Jasser A.M (2006). Extended – Spectrum Beta – Lactamases (ESBLs): A Global Problem. Kuwait Medical Journal, 38(3):171-185.
Bisht R., Katiyar A., Singh R and Mittal P (2009). Antibiotic Resistance – A Global Issue of Concern. Asian Journal of Pharmaceutical and Clinical Research, 2 (2):34-39.
Block S.S (2001). Disinfection, sterilization and preservation. 5th edition. Lippincott Williams & Wilkins, Philadelphia and London.
Cars O and Nordberg P (2005). Antibiotic resistance: The faceless threat. International Journal of Risk & Safety in Medicine, 17 (3/4): 103-110.
Ejikeugwu Chika, Iroha Ifeanyichukwu, Adikwu Michael and Esimone Charles (2013). Susceptibility and Detection of Extended Spectrum β-Lactamase Enzymes from Otitis Media Pathogens. American Journal of Infectious Diseases. 9(1):24-29.
Finch R.G, Greenwood D, Norrby R and Whitley R (2002). Antibiotic and chemotherapy, 8th edition. Churchill Livingstone, London and Edinburg.
Joslyn, L. J. (2000). Sterilization by Heat. In S. S. Block (Ed.), Disinfection, Sterilization, and Preservation (5th ed., pp. 695-728). Philadelphia, USA: Lippincott Williams and Wilkins.
Lai P.K and Roy J (2004). Antimicrobial and chemopreventive properties of herbs and spices. Curr. Med. Chem, 11 (11): 1451–1460.
Livermore D.M (2004). The need for new antibiotics. Clinical Microbiology & Infection, 4(10): 1-9.
Mascaretti O.A (2003). Bacteria versus antibacterial agents: An integrated approach. Washington: ASM Press.
Nally J.D (Ed.) (2007). Good manufacturing practices for pharmaceuticals. Sixth edition. Informa Healthcare USA, Inc, New York.
Discover more from Microbiology Class
Subscribe to get the latest posts sent to your email.