Metallo-beta-lactamases (MBLs) are beta-lactamase enzymes produced by pathogenic bacteria, and which hydrolyzes the carbapenems (e.g. imipenem, meropenem, and ertapenem) and render the antibiotics ineffective for treatment. They are encoded by genes that have been procured by pathogenic bacteria either by mutation or by horizontal gene transfer from other resistant microbes. MBLs efficiently hydrolyze all beta-lactam drugs except aztreonam, a monobactam. However, they confer variable range of high levels of resistance to all beta-lactam antibiotics and some non-beta-lactams such as fluoroquinolones and aminoglycosides; and their presence in clinically important Gram negative bacteria have put the use of the carbapenems under threat. MBL enzymes have high affinity for zinc ions (Zn2+); and thus the enzyme is largely inhibited by chelating agents (e.g. EDTA and dipicolinic acid) in vitro.
The susceptibility of MBL-producing bacteria to chelating agents is the basis upon which the enzyme can be detected phenotypically in pathogenic microorganisms. Genes responsible for the expression of MBLs, multidrug-hydrolyzing enzymes can also be chromosomally or plasmid-mediated as is also applicable with extended spectrum beta-lactamases (ESBLs) and other resistance genes harboured by pathogenic microorganisms. The carbapenems as earlier listed are often the last line of treatment option for a variety of infectious disease including those caused by multidrug resistant organisms (e.g. ESBL positive bacteria). The carbapenems are very potent antimicrobial agents used for the treatment of serious Gram negative bacterial infections, and because of the broad spectrum activity and stability of the carbapenems to most beta-lactamase enzymes, they have been widely used under restricted conditions in most hospitals worldwide as the first-line treatment for severe Gram negative infections.
MBLS are mainly expressed by non-lactose fermenters such as the Pseudomonas species and Acinetobacter species. The Enterobacteriaceae including Escherichia coli and Klebsiella species have also been reported to express MBLs. The presence or occurrence of MBL-producing bacteria in a hospital setting poses not only a therapeutic problem but also serious concern for infection control management in the health system. This is due to the fact that organisms producing MBLs are multidrug resistant in nature; and they limit therapeutic options for treatment. MBLs may be disseminated in hospital environment through genetic transfer elements such as transposons, plasmids and integrons amongst clinically important bacteria; and the extensive use of the carbapenems especially irrationally has given impetus to the spread of organisms that produce the enzymes. Pathogenic bacteria producing metallo-beta-lactamases (MBLs) now occur worldwide, and hospital infections have been reported in Europe, Asia, and America and in Africa.
The detection of MBL-producing bacteria is usually two-folds and involves a phenotypic screening test (Figure 1) and confirmatory tests as well as the application of PCR technique to amplify MBL genes in clinically important bacterial isolates. The phenotypic detection methods for MBLs from clinically important Gram negative bacteria make use of tests such as disk approximation, disk diffusion, micro-dilution test, E-test, and carbapenem hydrolysis test – all of which are usually performed based on routine clinical microbiology antimicrobial susceptibility test with some modifications. Particularly, the susceptibility of the test pathogen to any of the carbapenems (e.g. imipenem) is first evaluated by the Kirby-Bauer disk diffusion test; and organisms found to be resistant to any of these agents (based on the breakpoints of the Clinical Laboratory Standard Institute, CLSI) is subjected to a phenotypic confirmation test for MBL detection.
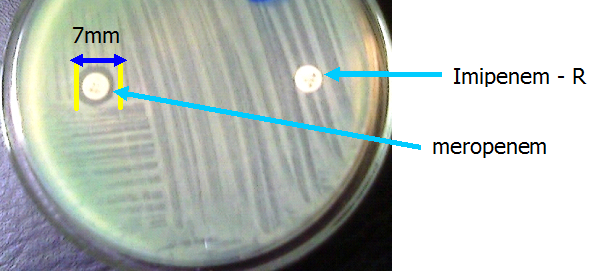
The disk potentiation or inhibitor-based assays (Figure 2) and the modified Hodges test (Figure 3) and are usually used for the phenotypic confirmation of MBL production in bacteria. In the inhibitor based test, MBL production in the test pathogen is detected phenotypically by using the carbapenem alone and in combination with a chelating agent (e.g. imipenem-EDTA). An increase in the zone diameter of any of the carbapenems when used alone and in combination with a chelating agent (i.e. imipenem-EDTA for example) infers MBL production phenotypically. Proper antimicrobial susceptibility testing (AST) is critical for the containment of MBL-producing bacteria, and AST will also help to guide therapy as well as prevent the emergence and spread of MBL genes amongst clinical and environmental isolates.
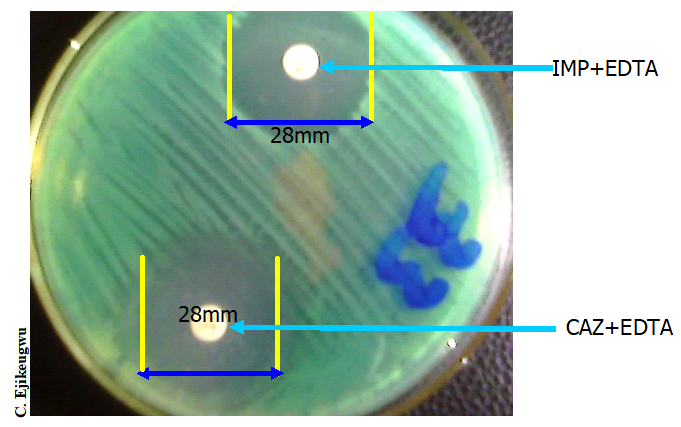

KEY: IPM-imipenem 10 µg, MEM-meropenem 10µg, EDTA (ethylenediamine tetraacetic acid).
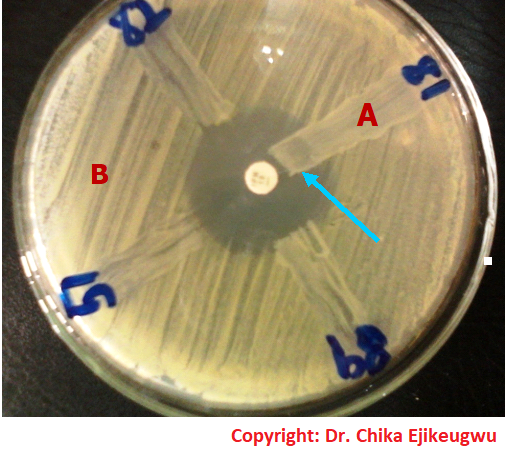

From Fig. 3: The test organism is E. coli strains (Nos. 82, 81, 89 and 51) producing metallo-β-lactamase (MBL) enzyme phenotypically. Strain No. 81 expressed MBL phenotypically, and this is noted by the inability of the test antibiotic (imipenem-10 µg) to inhibit the growth of the organism. MHT is performed by inoculating a Mueller-Hinton (MH) agar plate with a suspension of E. coli ATCC 25922 (adjusted to 0.5 McFarland turbidity standards) as per the CLSI criteria. A carbapenem disk (e.g. imipenem-10 10 µg) is aseptically placed at the center of the MH agar inoculated plate. A sterilized inoculating loop is used to streak the test bacteria (from an overnight culture) onto the MH agar plate in a straight line as shown in this illustration. The test organism should be heavily streaked (i.e. about 3-5 colonies should be picked), and the presence of growth of the test bacteria towards the carbapenem disk (arrows) is indicative of MBL production phenotypically. The test result is reported as positive in this case.
References
Arora D.R (2004). Quality assurance in microbiology. Indian J Med Microbiol, 22:81-86.
Ashutosh Kar (2008). Pharmaceutical Microbiology, 1st edition. New Age International Publishers: New Delhi, India.
Axelsen P.H (2002). Essentials of antimicrobial pharmacology. Humana Press, Totowa, New Jersey, USA. Al-Jasser A.M (2006). Extended – Spectrum Beta – Lactamases (ESBLs): A Global Problem. Kuwait Medical Journal, 38(3):171-185.
Bisht R., Katiyar A., Singh R and Mittal P (2009). Antibiotic Resistance – A Global Issue of Concern. Asian Journal of Pharmaceutical and Clinic Research, 2 (2):34-39.
Block S.S (2001). Disinfection, sterilization and preservation. 5th edition. Lippincott Williams & Wilkins, Philadelphia and London.
Cars O and Nordberg P (2005). Antibiotic resistance: The faceless threat. International Journal of Risk & Safety in Medicine, 17 (3/4): 103-110.
Carson C.F., Hammer K.A and Riley T.V (2006). Malaleuca alternifolia (Tea Tree) oil: A Review of Antimicrobial and other Medicinal Properties. Clinical Microbiology Review, 19(1):50-62.
Cowan M.M (1999). Plant products as antimicrobial agents. Clinical Microbiology Reviews., 564-582.
Denyer S.P., Hodges N.A and Gorman S.P (2004). Pharmaceutical Microbiology. 7th ed. Blackwell Publishing Company, USA.
Ejikeugwu Chika, Ikegbunam Moses, Ugwu Chigozie, Eze Peter, Iroha Ifeanyichukwu, and Esimone Charles (2013). Phenotypic Detection of Klebsiella pneumoniae Strains – Producing Extended Spectrum β-Lactamase (ESBL) Enzymes. Scholars Academic Journal of Biosciences. 1(1):20-23.
Ejikeugwu Chika, Iroha Ifeanyichukwu, Adikwu Michael and Esimone Charles (2013). Susceptibility and Detection of Extended Spectrum β-Lactamase Enzymes from Otitis Media Pathogens. American Journal of Infectious Diseases. 9(1):24-29.
Ejikeugwu Chika, Iroha Ifeanyichukwu, Adikwu Michael and Esimone Charles (2013). Susceptibility and Detection of Extended Spectrum β-Lactamase Enzymes from Otitis Media Pathogens. American Journal of Infectious Diseases. 9(1):24-29.
Finch R.G, Greenwood D, Norrby R and Whitley R (2002). Antibiotic and chemotherapy, 8th edition. Churchill Livingstone, London and Edinburg.
Joslyn, L. J. (2000). Sterilization by Heat. In S. S. Block (Ed.), Disinfection, Sterilization, and Preservation (5th ed., pp. 695-728). Philadelphia, USA: Lippincott Williams and Wilkins.
Lai P.K and Roy J (2004). Antimicrobial and chemopreventive properties of herbs and spices. Curr. Med. Chem, 11 (11): 1451–1460.
Livermore D.M (2004). The need for new antibiotics. Clinical Microbiology & Infection, 4(10): 1-9.
Mascaretti O.A (2003). Bacteria versus antibacterial agents: An integrated approach. Washington: ASM Press.
Nally J.D (Ed.) (2007). Good manufacturing practices for pharmaceuticals. Sixth edition. Informa Healthcare USA, Inc, New York.
Discover more from #1 Microbiology Resource Hub
Subscribe to get the latest posts to your email.


