Genomic DNA is a double helix structure that is composed of several components including purines (adenine and guanine) and pyrimidines (cytosine and thymine) as nitrogenous bases, deoxyribose sugar (a pentose sugar) and phosphate groups. It is a large nucleic acid molecule composed mainly of two polynucleotide chains or strands that are coiled in a complementary fashion to form a double helix structure.
Each of the two polynucleotide chains that make up the double helix structure of genomic DNA are complementary in nature because the purine and pyrimidines are linked in a base-pairing fashion that always ensures that a particular purine is constantly linked to a particular pyrimidine at each stage of the double helix structure.
For example, adenine (A) always pairs with thymine (T) while guanine (G) always pairs with cytosine (C) and vice versa. The double helix structure of DNA is held together by strong covalent or hydrogen bonds mainly phosphodiester bonds. Adenine is paired to thymine by two hydrogen bonds while guanine is paired to cytosine by three hydrogen bonds (Figure 1).
It is noteworthy that in ribonucleic acid (RNA), thymine is replaced by uracil (U), and the pentose sugar is known as ribose sugar. RNA is single-stranded unlike DNA which is double stranded. The base-pairing between the purines and pyrimidines in the double helix structure of the DNA shows the complementarity nature of the genomic DNA, which is the main genetic material of the cell.
Genomic DNA is found in both prokaryotic and eukaryotic cells and the genetic information it encodes is transcribed and translated into protein molecules with unique biological functions in the cell of an organism. On close inspection of the DNA double helix structure, the base-pairing pattern of the nitrogenous bases (i.e. the purines and pyrimidines) with the phosphate groups makes the genomic DNA to resemble a ladder. While one end of the double helix structure of the genomic DNA has a free 3ꞌ hydroxyl (OH–) group, the other end has an exposed 5ꞌ OH– group. DNA replication always proceeds in the 5ꞌ to 3ꞌ direction and not the other way round.
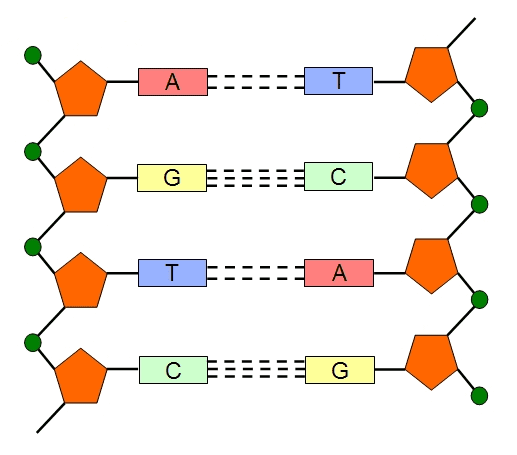
Three hydrogen bonds hold guanine to cytosine while two hydrogen bonds hold adenine to thymine. The G+C content of a DNA molecule is often determined from the melting temperature (Tm) of the nucleic acid (i.e. DNA); and this is because DNA molecules with higher content of guanine and cytosine (i.e. G+C content) have greater amount of hydrogen bonds and thus such double stranded DNA molecules will require a higher temperature for its separation into single strands. Therefore, DNA molecules with higher G+C content often have higher melting point. The green colour represents the phosphate backbone of the DNA while the yellow colour represents the pentose sugar (i.e. deoxyribose sugar).
Genomic DNA in bacteria (e.g. Escherichia coli) is circular in structure or shape except in some prokaryotic cells (e.g. Borrelia species) whose DNA is linear like that of the eukaryotes. However, the genomic DNA of some microorganisms especially those of eukaryotic organisms are linear in nature. Thus, genomic DNA can either be linear or circular in shape. The genomic DNA in both prokaryotic and eukaryotic cells is usually complexed with specialized proteins such as histones in eukaryotes to form the chromosomes.
References
Alberts B, Bray D, Lewis J, Raff M, Roberts K and Watson J.D (2002). The molecular Biology of the Cell. Fourth edition. New York, Garland, USA.
Chen I and Dubnau D (2004). DNA uptake during bacterial transformation. Nat. Rev. Microbiol. 2 (3): 241–249.
Cooper G.M and Hausman R.E (2004). The cell: A Molecular Approach. Third edition. ASM Press.
Dale J (2003). Molecular genetics of bacteria. Jeremy W. Dale and Simon Park (4th eds.). John Wiley & Sons Ltd, West Sussex, UK. Pp. 312-313.
Das H.K (2010). Textbook of Biotechnology. Fourth edition. Wiley edition. Wiley India Pvt, Ltd, New Delhi, India.
Lewis R (2004). Human Genetics: Concepts and Applications. Sixth edition. McGraw Hill Publishers, USA.
Lodish H, Berk A, Matsudaira P, Kaiser C.A, Kreiger M, Scott M.P, Zipursky S.L and Darnell J (2004). Molecular Cell Biology. Fifth edition. Scientific American Books, Freeman, New York, USA.
Madigan M.T., Martinko J.M., Dunlap P.V and Clark D.P (2009). Brock Biology of Microorganisms, 12th edition. Pearson Benjamin Cummings Inc, USA.
McPherson M and Moller S (2002). PCR: The Basics. 2nd edition. Taylor and Francis Group. New York, USA.
Sambrook, J., Russell, D.W. (2001). Molecular Cloning: a Laboratory Manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York.
Synder L, Peters J.E, Henkin T.M and Champness W (2013). Molecular Genetics of Bacteria. Fourth edition. American Society of Microbiology Press, USA.
Tamarin Robert H (2002). Principles of Genetics. Seventh edition. Tata McGraw-Hill Publishing Co Ltd, Delhi.
Discover more from #1 Microbiology Resource Hub
Subscribe to get the latest posts to your email.


