Genes in the DNA code for proteins; and it is the gene that directs the cell in what particular order to assemble the amino acids which eventually becomes the building blocks of protein molecules. The cell of an organism must use one nucleotide or more in the DNA to spell out or specify each of the amino acid in a particular protein; and this is because there are 20 possible essential amino acids and only four (4) possible bases (i.e. guanine-G, adenine-A, thymine-T and cytosine-C) that are mainly involved in the protein synthesis process as well as in other genetic processes. The 20 essential amino acids are shown in Table 1.
Each possible set of three nucleotides (i.e. a codon) in the DNA is what specifies one particular amino acid, which in combination with other amino acids forms a protein. For example, AAA specifies the amino acid lysine while GCT specifies alanine. A codon is a triplet of three nucleotides in the mRNA that codes for a specific amino acid during the synthesis of a protein molecule. Itis a sequence of nucleotides formed by triplet of bases (e.g. TAC is the codon for the amino acid tyrosine).
Table 1. The Twenty Essential Amino Acids
| Amino acid | Abbreviation |
| Alanine | ALA |
| Arginine | ARG |
| Asparagines | ASN |
| Aspartic acid | ASP |
| Cysteine | CYS |
| Glutamine | GLN |
| Glutamic acid | GLU |
| Glycine | GLY |
| Histidine | HIS |
| Isoleucine | ILE |
| Leucine | LEU |
| Lysine | LYS |
| Methionine | MET |
| Phenylalanine | PHE |
| Proline | PRO |
| Serine | SER |
| Threonine | THR |
| Tryptophan | TRP |
| Tyrosine | TYR |
| Valine | VAL |
Codon provides the genetic information which causes a specific amino acid to be produced in a cell. There are various types of codons, and each performs a specific function during protein synthesis in the cell of an organism. A start codon is the codon that signals translation, and it tells the mRNA when to initiate the peptide chain formation or elongation. It is the first codon of an mRNA transcript that is translated by the ribosomes during protein synthesis; and start codons are also known as initiator codons because they signal and initiate the translation process. AUG is a start codon, and it also codes for the amino acid, methionine.
Apart from signaling the incorporation of the amino acid methionine into the growing polypeptide chain, the start codon AUG also signals the start of translation during protein synthesis in the cell. Stop codons are triplet of nucleotide sequence that signals the end of translation. They are also known as termination or nonsense codons because they bring protein synthesis in the cell to an end. Examples of stop codons include UAG, UAA, and UGA.
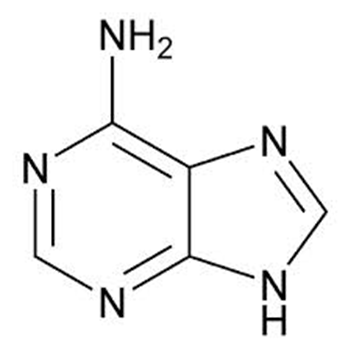
In the genetic code, there are 64 possible codons; and this comprises 3 nucleotides in each codon with 4 possible bases which are adenine (Figure 1). Adenine is a purine.
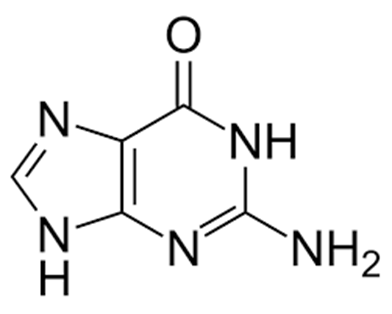
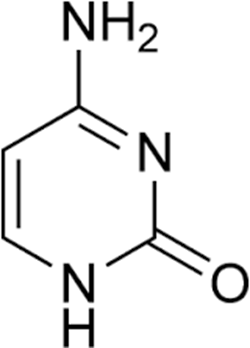
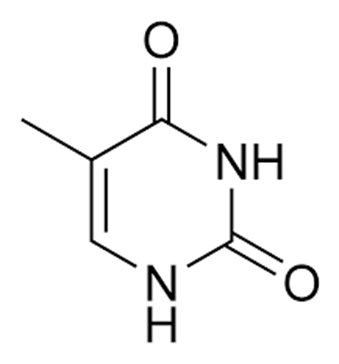
Guanine (Figure 2), a purine is another type of nitrogenous base. Cytosine (Figure 3) and thymine (Figure 4) which are both pyrimidines are other examples of nitrogenous bases that are found in DNA molecules.
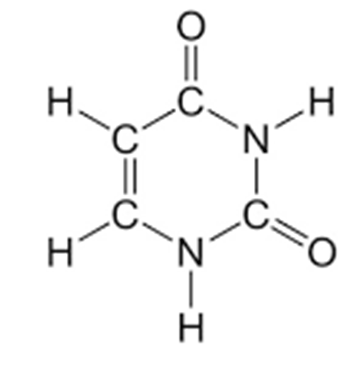
Uracil (Figure 5), a pyrimidine is another type of nitrogenous base which is only found in RNA molecules alone, where it replaces thymine (T). The language of the gene is created when these bases (i.e. A, G, C, and T) are genetically arranged or set in the DNA just the same way the 26 English alphabets are ordered to create meaningful words.
These four bases which includes adenine (A), guanine (G), cytosine (C), and thymine (T), and which are usually found in the mRNA of the cell can generate 64 possible triplet combinations (e.g. UAG). Out of this 64 possible triplet combinations or codons, 61 triplet combinations encode the 20 essential amino acids (Table 1) while the remaining three triplet combinations (UAG, UGA & UAA) are generally known as stop or nonsense codons because they signify the termination of the polypeptide chain formation during protein synthesis in the ribosome.
Mathematically, 43 = 64 possible codons; where “4” refer to the bases (i.e. A, G, C & T)and “3” refer to the triplet combinations or codons.
References
Alberts B, Bray D, Johnson A, Lewis J, Raff M, Roberts K and Walter P (1998). Essential Cell Biology: An Introduction to the Molecular Biology of the Cell. Third edition. Garland Publishing Inc., New York.
Dale J (2003). Molecular genetics of bacteria. Jeremy W. Dale and Simon Park (4th eds.). John Wiley & Sons Ltd, West Sussex, UK.
Edelstein, M. L., Abedi, M. R., Wixon, J., and Edelstein, R. M. (2004). Gene therapy clinical trials worldwide 1989-2004—an overview. J Gene Med, 6: 597-602.
Ferrua, F.; Brigida, I.; Aiuti, A. (2010). Update on gene therapy for adenosine deaminase-deficient severe combined immunodeficiency. Current Opinion in Allergy and Clinical Immunology. 10 (6): 551–556.
Gardlík R, Pálffy R, Hodosy J, Lukács J, Turna J, Celec P; Pálffy; Hodosy; Lukács; Turna; Celec (2005). Vectors and delivery systems in gene therapy. Med Sci Monit. 11 (4): RA110–21.
Horn PA, Morris JC, Neff T, Kiem HP; Morris; Neff; Kiem (2004). Stem cell gene transfer—efficacy and safety in large animal studies. Mol. Ther. 10 (3): 417–31.
S Li and L Huang (2000). Nonviral gene therapy: promises and challenges. Gene Therapy, 7:31-34. www.nature.com/gt
Discover more from Microbiology Class
Subscribe to get the latest posts sent to your email.