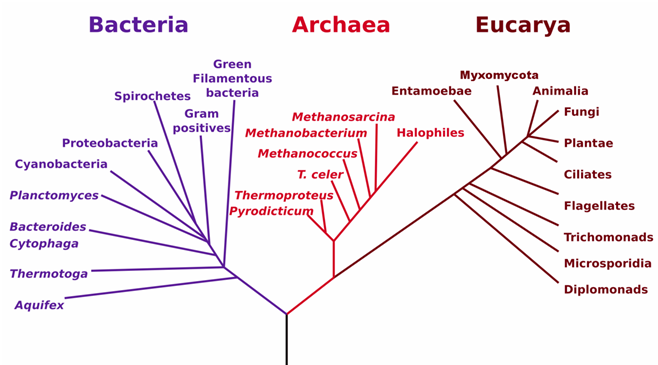
Archaea bacteria generally inhabit terrestrial and aquatic environments where the condition of living is extremely harsh and unfavourable for other microbial cells. Habitats occupied by Archaea have very high temperature and salinity. Archaea are known to live in hostile environments including hot springs and areas with high salt concentration. They are not known to cause any disease as their prokaryotic relatives, bacteria. Both Archaea and bacteria differ greatly in their morphological structures, physiology and metabolic activities. Various forms of the Archaea bacteria are known to exist and they include the hyper or extreme thermophiles, the methanogens, the extreme halophiles and the sulphate-reducing Archaea (e.g. Archaeoglobus species). Some Archaea bacteria such as the extreme or hyper-thermophilc organisms (e.g. Sulfolobus acidocaldarius) are highly adapted to living at very high temperature above 100oC.
Thermus aquaticus is an Archaea but not an extreme thermophile; instead T. aquaticus isa thermophile or thermophilic bacterium that lives in habitats with temperature below 100oC. T. aquaticus is the source of the enzyme Taq polymerase which is used in the polymerase chain reaction (PCR), animportant laboratory technique in molecular biology. Other examples of thermophilic Archaea bacteria include Thermoplasma species, bacteria without cell walls. Methanogens such are Archaea that produce methane (CH4) gas; and they also occur in the stomach of ruminant animals (e.g. cow). The methanogens that exist in the stomach of animals are known as animal methanogens while those that occur in the natural environment i.e. in harsh terrestrial and aquatic habitats are generally referred to as symbiotic methanogens. The metabolic activities of methanogens are responsible for the world’s reserve of natural gases which are tapped all over the globe for various industrial and economic purposes. Examples of methanogens include Haloferax volcanni and Methanosarcina barkeri.
The extreme halophiles are Archaea that live in habitats with high salt concentration. Extreme halophiles will not grow at low salt concentrations because they are organisms that require high salt concentration for growth. Examples of halophiles include Halobacterium salinarium and Halobacterium halobium. Archaea bacteria differ greatly from eukaryotic cells and bacteria; and they only exist in extreme and harsh environments due to specialized structures as well as their metabolic adaptations which allow them to inhabit such environments. They are mostly Gram-negative anaerobic bacteria but some are Gram-positive (e.g. Methanobacterium formicicum); and Archaea bacteria differ from other organisms in their physiology, morphology, ecology and reproduction.
Eukarya domain mainly contains all eukaryotic microorganisms such as fungi and algae; and plants, lichens and animals which are known as macro-organisms. Slime moulds and diatoms as well as other protists or protozoa and ciliates are also in the Eukarya domain. The Eukarya domain is unique and it contains several eukaryotic microorganisms (i.e. cells with membrane-bound nucleus and other membrane-bound organelles) that are distinct from the Archaea and Bacteria domains. They also contain organisms that are pathogenic in man and animals; and several of the eukaryotic microorganisms and macroorganisms are non-pathogenic and beneficial to man and his environment. Eukaryotic cells are distinct from prokaryotic cells- which do not have a membrane-bound nucleus as well as other membrane bound organelles.
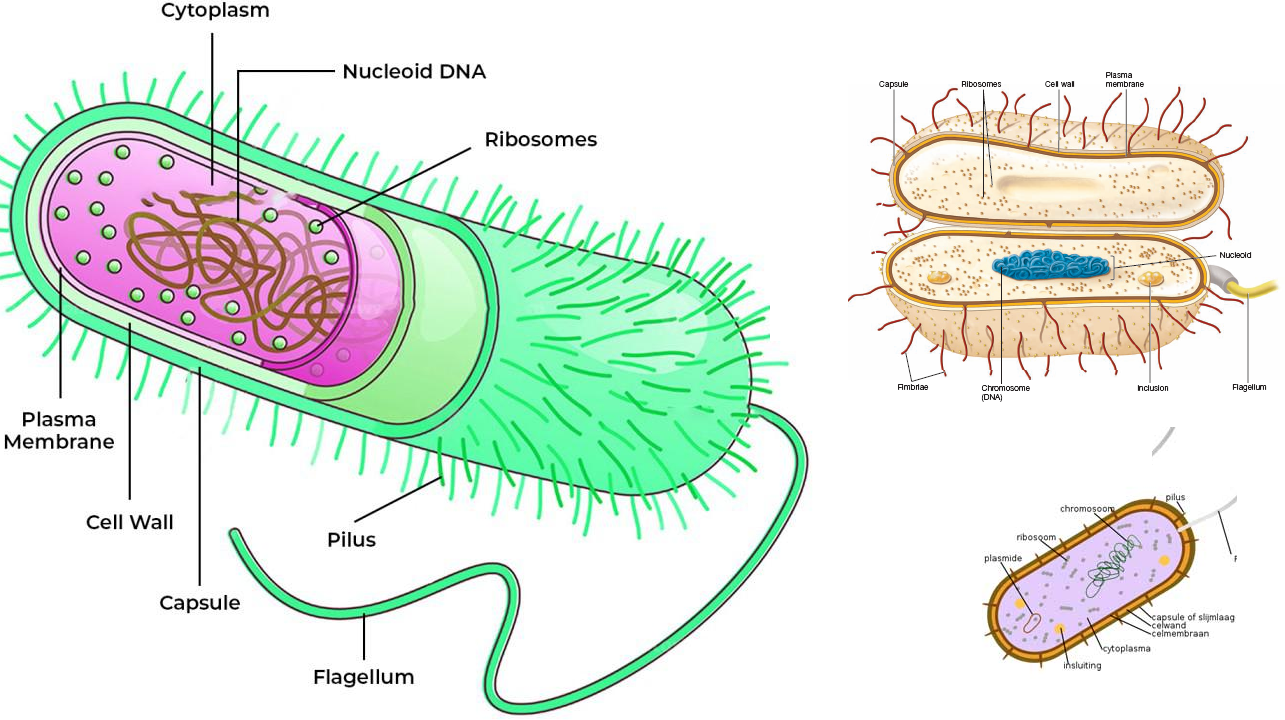
Bacteria are microbial cells that are neither members of the Eukarya or Archaea domain, but they are basically prokaryotic cells (i.e. organisms without a true nucleus). They are all prokaryotic organisms, and are ubiquitously found in the environment. Bacteria domain includes both pathogenic and beneficial or non-pathogenic bacteria species. Some recognized major phyla of the Bacteria domain which represents different genera of bacteria include Proteobacteria, Green filamentous bacteria, Spirochaetes, Actinobacteria, Fusobacteria, Cyanobacteria, Acidobacteria, Verrucomicrobia, Planctomycetes, Aquifex, Chlamydia and Deinococcus phylum (which contains D. radiodurans– that survives high radiation) et cetera. The Proteobacteria is the largest phylum of the Bacteria domain; and this division contains several genera of bacteria including chemolithotrophic bacteria, phototrophic bacteria as well as numerous species of bacteria that are pathogenic and non-pathogenic in nature.
Five (5) classes of the proteobacteria phyla are known, and these are: alpha (α) proteobacteria (e.g. Rhizobium), beta (β) proteobacteria (e.g. Neisseria), gamma (γ) proteobacteria (e.g. Escherichia), delta (δ) proteobacteria (e.g. Myxococcus) and epsilon (ε) proteobacteria (e.g. Helicobacter). Chemolithotrophs are organisms (Archaea and bacteria inclusive) that obtain their energy from the oxidation of inorganic compounds, and such cells are said to be chemolithotrophic in their mode of nutrition. Phototrophs or phototrophic bacteria such as cyanobacteria and green bacteria obtain their energy through the process of photosynthesis, like plants. Based on their cell walls, bacteria are classified as either Gram-positive or Gram-negative. Some bacteria are neither Gram-positive nor Gram-negative, and are generally known as Gram variable bacteria. In terms of oxygen requirement, the organisms in the Bacteria domain are aerobic, anaerobic, micro-aerobic or facultative aerobes or anaerobes.
References
Ciccarelli FD, Doerks T, von Mering C, Creevey CJ, Snel B, Bork P (2006). Toward automatic reconstruction of a highly resolved tree of life. Science, 311(5765):1283–1287.
Euzéby, J. P. (2010). List of Bacterial Names with Standing in Nomenclature. Int. J. Syst. Bacteriol., 47, 8 July, 2016. Retrieved from http://www.bacterio.cict.fr/m/micrococcus.html
Fauquet C.M, Mayo M.A, Maniloff J, Desselberger U and Ball L.A. (Eds) (2005). Virus Taxonomy. Eight Report of the International Committee on Taxonomy of Viruses. Burlington, M.A. Elsevier Academic Press.
George M. Garrity (2005). Bergey’s manual of systematic bacteriology. 2. Auflage. Springer, New York, 2005, Volume 2: The Proteobacteria, Part B: The Gammaproteobacteria.
Grenfell B.T, Pybus O.G, Gog J.R, Wood J.L, Daly J.M, Mumford J.A and Holmes E.C (2004). Unifying the Epidemiological and Evolutionary Dynamics of Pathogens. Science, 303(5656): 327–332.
Gupta RS (2000). The natural evolutionary relationships among prokaryotes”. Crit. Rev. Microbiol, 26 (2):111–131.
Janda, J. M., and Abbott, S. L. (2006). The Genera Klebsiella and Raoultella. The Enterobacteria (2nd ed., pp. 115-129). Washington, USA: ASM Press.
Madigan M.T., Martinko J.M., Dunlap P.V and Clark D.P (2009). Brock Biology of microorganisms. 12th edition. Pearson Benjamin Cummings Publishers. USA. Pp.795-796.
Moreira D and López-García P (2009). Ten reasons to exclude viruses from the tree of life Nature Reviews Microbiology, 7(4):306-311.
Murray P.R, Baron E.J, Jorgensen J.H., Pfaller M.A and Yolken R.H (2003). Manual of Clinical Microbiology. 8th edition. Volume 1. American Society of Microbiology (ASM) Press, Washington, D.C, U.S.A.
Purves W.K, Sadava D, Orians G.H, and Heller H.C (2007). Life: The Science of Biology, 7e by Sinauer Associates, W.H. Freeman and Company, Sunderland.
Salyers A.A and Whitt D.D (2001). Microbiology: diversity, disease, and the environment. Fitzgerald Science Press Inc. Maryland, USA.
Woese C.R, Kandler O, Wheelis M.L (1990). Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proceedings of the National Academy of Science, 87(12):4576–4579.
Discover more from Microbiology Class
Subscribe to get the latest posts sent to your email.