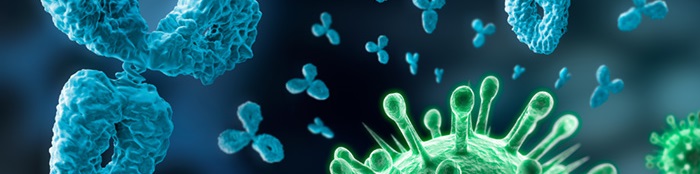
Viruses are infectious agents that have a simple acellular structure that is mainly made up of a protein coat or capsid and a nucleic acid genome which can either be DNA or RNA. Some viruses also have envelopes (which are lipid-containing outer membranous layer that surround the nucleocapsid in some viruses) while others lack them, and are thus generally known as naked viruses.
Viruses with envelopes are known as enveloped viruses. Viruses are unique group of microorganisms that are composed of several chemical molecules or structures that are vital to their development and replication within a particular host cell. These chemical compositions of viruses include the viral proteins, viral envelopes, viral glycoproteins and viral genomes; and they form the integral parts of mature viruses.
- Viral proteins: Viral proteins are fundamental structural and functional components of viruses, playing central roles in infectivity, host interaction, immune recognition, and viral stability. They constitute the primary antigenic determinants (epitopes) of an infecting virus and are therefore the principal targets of host immune responses, including neutralizing antibodies and T-cell–mediated immunity. The antigenic properties of viral proteins underpin serological diagnostics and vaccine design, as protective immunity is largely directed against specific surface-exposed viral proteins. A critical function of viral proteins is mediating attachment to and entry into host cells. Specialized viral surface proteins—often referred to as attachment proteins, glycoproteins, or spikes—bind with high specificity to receptor molecules on the surface of susceptible host cells. This receptor–ligand interaction determines host range, tissue tropism, and species specificity. Following receptor binding, viral proteins facilitate membrane fusion, endocytosis, or other entry mechanisms that enable penetration of the viral particle or delivery of the viral genome into the host cytoplasm or nucleus. In many enveloped viruses, conformational changes in viral glycoproteins trigger fusion between viral and cellular membranes, while in non-enveloped viruses, capsid proteins mediate endosomal escape. Structurally, viral proteins form the capsid, also known as the protein coat, which encloses and protects the viral nucleic acid genome (DNA or RNA). Capsid proteins assemble into highly ordered structures exhibiting defined symmetry, commonly icosahedral or helical. This structural organization ensures efficient genome packaging while maintaining stability and infectivity. The capsid serves as a protective barrier against physical, chemical, and enzymatic damage, including degradation by host nucleases that would otherwise hydrolyze the viral genome. In enveloped viruses, internal structural proteins (e.g., matrix proteins) further stabilize the virion by linking the nucleocapsid to the lipid envelope. Beyond structural roles, many viral proteins are non-structural and are synthesized only during infection. These proteins regulate viral genome replication, transcription, protein processing, and assembly. Some viral proteins modulate host cellular pathways, suppress innate immune responses, and manipulate host machinery to favor viral replication. Viral proteins provide structural integrity, protect genomic material, mediate host cell entry, and orchestrate replication processes. Their multifunctional roles make them indispensable to viral survival, pathogenicity, and transmission.
- Viral enzymes: Viral enzymes are specialized proteins encoded by viruses to facilitate essential steps in their replication and propagation within host cells. Although viruses rely heavily on host cellular machinery for energy production and many biosynthetic functions, numerous viruses carry or encode their own enzymes to ensure efficient genome replication, transcription, integration, and maturation. These enzymes are proteinaceous in nature and are often packaged within the virion or synthesized immediately after infection. A prominent example is reverse transcriptase (RT), an RNA-dependent DNA polymerase found in retroviruses such as Human immunodeficiency virus 1. RT catalyzes the synthesis of complementary DNA (cDNA) from a viral RNA template, a critical step that enables integration of the viral genome into the host’s chromosomal DNA. This integration, mediated by another viral enzyme called integrase, allows the virus to exploit host transcriptional machinery for the production of viral proteins and progeny genomes. Without reverse transcriptase, retroviral replication would not be possible. Similarly, many RNA viruses encode RNA-dependent RNA polymerase (RdRp), an enzyme that synthesizes RNA from an RNA template. This enzyme is essential because host cells lack polymerases capable of replicating RNA from RNA templates. For example, viruses such as SARS-CoV-2 depend on RdRp to replicate their positive-sense RNA genomes and to generate subgenomic RNAs required for protein synthesis. While these enzymes are indispensable for viral replication, they generally do not contribute structurally to the viral capsid or protein coat. Instead, their function is catalytic and regulatory, ensuring that viral genetic material is accurately replicated and expressed once inside the host cell. Thus, viral enzymes represent critical molecular tools that enable viruses to complete their life cycles and are key targets for antiviral drug development.
- Viral genomes: The viral genome is the fundamental repository of genetic information required for the replication and propagation of a virus within a susceptible host cell. Unlike cellular organisms, which universally utilize double-stranded DNA as their genetic material, viruses exhibit remarkable diversity in genome composition and organization. A defining principle in virology is that a virus contains either DNA or RNA as its nucleic acid genome, never both simultaneously. This binary distinction forms the basis of major viral classification systems and profoundly influences replication strategy, mutation rate, and evolutionary dynamics. Viral genomes occur in multiple structural configurations. They may be linear or circular, and can be single-stranded (ss) or double-stranded (ds). These structural attributes are not trivial; they determine how the genome is replicated, transcribed, and packaged. For example, double-stranded genomes resemble cellular nucleic acids more closely and typically rely on DNA-dependent polymerases, whereas single-stranded genomes require synthesis of a complementary strand before transcription or replication can proceed. Another important genomic feature of the viral genome is segmentation. Viral genomes may be segmented, meaning the genetic material is divided into two or more distinct nucleic acid molecules, or non-segmented, consisting of a single continuous molecule. Segmented genomes permit reassortment – the exchange of genome segments between related viruses infecting the same cell which can generate substantial genetic diversity. In contrast, non-segmented genomes evolve primarily through mutation and recombination. Among RNA viruses, genome polarity-or sense is a critical determinant of replication strategy. Positive-sense (+) single-stranded RNA viruses possess genomes that function directly as messenger RNA (mRNA). Upon entry into the host cell, their genomic RNA can be immediately translated by host ribosomes to produce viral proteins. In contrast, negative-sense (−) single-stranded RNA viruses carry genomes complementary to mRNA. These viruses must package an RNA-dependent RNA polymerase within the virion to synthesize a positive-sense intermediate before translation can occur. A distinct category within RNA viruses is the ambisense genome organization, in which portions of the RNA genome encode proteins in both positive-sense and negative-sense orientations. This arrangement requires regulated transcription from both strands during infection. Members of the Arenaviridae exemplify this ambisense strategy, illustrating the structural and functional flexibility of RNA viral genomes. The biochemical nature of viral genomes differs substantially between DNA and RNA viruses. DNA viruses generally possess genomes ranging from approximately 3 to 370 kilobase pairs (kbp). These genomes tend to be more genetically stable due to proofreading activity associated with many DNA polymerases. In contrast, RNA viruses typically have smaller genomes, approximately 7 to 30 kilobases (kb). The limited genome size of RNA viruses reflects the higher error rates of RNA-dependent RNA polymerases, which lack robust proofreading mechanisms. Elevated mutation rates contribute to rapid viral evolution, antigenic variation, and adaptation to selective pressures such as host immunity or antiviral drugs. In animal virology for example, several general patterns are observed. Most DNA viruses that infect animals and humans possess double-stranded DNA (dsDNA) genomes. A notable exception is the family Parvoviridae, whose members contain single-stranded DNA (ssDNA) genomes. Conversely, the majority of RNA viruses infecting animals and humans have single-stranded RNA (ssRNA) genomes. The principal exception is the family Reoviridae, which contains viruses with double-stranded RNA (dsRNA) genomes. The diversity in viral genome type, structure, segmentation, and polarity reflects evolutionary solutions to a common biological challenge: efficient replication within host cells while evading host defenses. Genome architecture dictates not only replication strategy but also pathogenic potential, host range, and mechanisms of genetic variation. Consequently, understanding viral genome organization is central to virology, informing diagnostics, antiviral drug development, vaccine design, and molecular epidemiology.
- Viral glycoproteins: Viral glycoproteins are complex macromolecules composed of a polypeptide backbone covalently linked to carbohydrate moieties through glycosylation. These molecules are encoded by viral genes and synthesized within infected host cells using the host’s translational and post-translational machinery. Glycosylation typically occurs in the endoplasmic reticulum and Golgi apparatus, where oligosaccharides are attached to specific amino acid residues, most commonly through N-linked or O-linked glycosidic bonds. The resulting glycoproteins are then transported to cellular membranes, where they become incorporated into newly forming virions. Viral glycoproteins are characteristic features of enveloped viruses. In these viruses, the glycoproteins are embedded within the lipid bilayer envelope, which is derived from host cellular membranes during viral budding. Prominent examples include the hemagglutinin and neuraminidase proteins of Influenza A virus and the spike (S) protein of SARS-CoV-2. These surface glycoproteins mediate viral attachment (adsorption) to specific receptors on host cells, determining host range and tissue tropism. Following receptor binding, many viral glycoproteins also facilitate membrane fusion, enabling entry of the viral genome into the host cytoplasm. Beyond their role in attachment and entry, viral glycoproteins are major antigenic determinants. They are primary targets of neutralizing antibodies and are therefore central to vaccine design and antiviral therapeutics. Glycosylation can shield critical epitopes from immune recognition, contributing to immune evasion. In contrast, non-enveloped (naked) viruses lack a lipid envelope and therefore do not possess viral glycoproteins embedded in a membrane. Instead, they rely on capsid proteins to mediate attachment and entry into host cells. Thus, viral glycoproteins are defining structural and functional components of enveloped viruses, essential for infectivity and host interaction.
- Viral envelopes: Viruses are commonly classified as either enveloped or non-enveloped (naked) based on the presence or absence of a surrounding lipid membrane. This structural distinction has major implications for viral entry, environmental stability, transmission dynamics, and susceptibility to disinfectants. Enveloped viruses possess an outer lipid bilayer known as the viral envelope. This envelope is not synthesized de novo by the virus; rather, it is acquired from the host cell during viral maturation. Specifically, as newly assembled nucleocapsids migrate to the host cell membrane, they bud through a cellular membrane—most commonly the plasma (cytoplasmic) membrane, though in some viruses the envelope may derive from internal membranes such as the endoplasmic reticulum or Golgi apparatus. During this budding process, the virus incorporates a portion of the host lipid bilayer into its structure. Embedded within this lipid bilayer are viral glycoproteins (spike proteins) that are encoded by the viral genome and inserted into the host membrane prior to budding. These glycoproteins are critical for host cell recognition, receptor binding, and membrane fusion during subsequent infection. Because the envelope is composed primarily of a lipid bilayer, enveloped viruses are relatively fragile in the external environment. They are highly susceptible to desiccation, heat, detergents, alcohols, ether, and other lipid-solvent organic agents. Disruption of the lipid envelope compromises the integrity of viral glycoproteins required for attachment and entry, rendering the virus non-infectious. For this reason, enveloped viruses typically require close contact, respiratory droplets, blood, or other bodily fluids for transmission. In contrast, naked (non-enveloped) viruses lack a lipid membrane and consist solely of a nucleic acid genome enclosed within a protein capsid. The absence of a lipid envelope confers greater structural stability. These viruses are generally resistant to lipid solvents, detergents, and environmental stressors such as drying and pH fluctuations. As a result, naked viruses often survive longer on surfaces and may be transmitted via the fecal–oral route or through contaminated fomites. The presence or absence of a viral envelope is a fundamental structural characteristic that influences viral stability, mechanisms of entry, modes of transmission, and susceptibility to chemical inactivation.
References
Acheson N.H (2011). Fundamentals of Molecular Virology. Second edition. John Wiley and Sons Limited, West Sussex, United Kingdom.
Alan J. Cann (2005). Principles of Molecular Virology. 4th edition. Elsevier Academic Press, Burlington, MA, USA.
Alberts B, Bray D, Johnson A, Lewis J, Raff M, Roberts K and Walter P (1998). Essential Cell Biology: An Introduction to the Molecular Biology of the Cell. Third edition. Garland Publishing Inc., New York.
Barrett J.T (1998). Microbiology and Immunology Concepts. Philadelphia, PA: Lippincott-Raven Publishers. USA.
Black, J.G. (2008). Microbiology: Principles and Explorations (7th ed.). Hoboken, NJ: J. Wiley & Sons.
Brian W.J Mahy and Mark H.C van Regenmortel (2010). Desk Encyclopedia of Human and Medical Virology. Elsevier Academic Press, San Diego, USA.
Brooks G.F., Butel J.S and Morse S.A (2004). Medical Microbiology, 23rd edition. McGraw Hill Publishers. USA.
Cann A.J (2011). Principles of Molecular Virology. Fifth edition. Academic Press, San Diego, United States.
Carter J and Saunders V (2013). Virology: Principles and Applications. Second edition. Wiley-Blackwell, New Jersey, United States.
Champoux J.J, Neidhardt F.C, Drew W.L and Plorde J.J (2004). Sherris Medical Microbiology: An Introduction to Infectious Diseases. 4th edition. McGraw Hill Companies Inc, USA.
Dimmock N (2015). Introduction to Modern Virology. Seventh edition. Wiley-Blackwell, New Jersey, United States.
Dimmock N.J, Easton A.J and Leppard K.N (2001). Introduction to modern virology. 5th edition. Blackwell Science publishers. Oxford, UK.
Discover more from Microbiology Class
Subscribe to get the latest posts sent to your email.